Viral gene delivery is adept at the persistent expression of therapeutic transgenes, e.g., for gene replacement therapies against rare diseases, or long-lived CAR-T cells. In contrast, nonviral genetic medicines can provide transient efficacy against conditions where doses must be safely repeated or “dialed-in.” In 1998, the late professor and gene therapy pioneer, Jon Wolff and Dr. Vladimir Trubetskoy said that therapeutic delivery of nonviral genetic material was in a “Cambrian” period of bustling trial-and-error evolution and innovation. [1] 4 years later, Dr. Wolff would dub it “The ‘grand’ problem of synthetic delivery.” [2]
Over two decades, we’ve seen some real successes. There’s mass-produced mRNA for two SARS-CoV-2 vaccines [3] and additionally, four siRNA drugs (ONPATTRO®, GIVLAARI®, LEQVIO®, OXLUMO™) [4] approved by regulatory authorities as of this writing. Yet today—while not an impossible challenge—efficient and non-cytotoxic delivery of nucleic acids sans viruses still remains a “grand problem.” [5] This is especially true for many primary cell types that lack a transformed phenotype. [6] Can this perennial pain point be remedied for ex vivo cell engineering of regenmed therapies? Here in this blog, we’ll show you one path forward to help navigate the pitfalls with mRNA transfection.
For many primary cells, nonviral gene delivery (NVGD) of nucleic acids like mRNA has two main issues: (1), low efficiency in percentage of robustly transfected cells; and (2), dead cells from cytotoxic and/or inflammasome effects via overloaded complexed nucleic acids. [7, 8, 9] Excess mRNA+lipoplexes does not yield happy cells, and thus each protocol needs a cell-specific optimization. But please indulge one more analogy from biology; we find that transfection is like farming. Let the “seeds” be the complexed mRNA and imagine the “crops” as transfected cells. Plenty of great work is already published on parameters of both the nucleic acid system [8, 10, 11, 12, 13, 14] as well as the cells [15, 16, 17] for nonviral delivery, but what about the “soil?” Why, that’s the media, of course!
On our “farm” (aka RoosterBio), we unleashed in 2021 a complete medium, RoosterGEM™, to assist in lentivirus-mediated transduction of human mesenchymal stromal/stem cells (hMSCs). When RoosterGEM’s “cell-grow-culture” conditions the hMSCs like “soil,” these cells are much more likely to swallow virus “cap-seeds” and express their synthetic genes. This bump in efficiency allows lower MOIs to enable the investigator to draw from costly lentivirus stocks more slowly—or to leverage them across a broader range of experimental variables for richer data. We’ve previously published blogs, webinars, and posters at national meetings on this new biotool. [18, 19, 20]
Fast forward to 2022. We’re quickly learning from ourselves and customers that the RoosterGEM “soil” works with a more diverse assortment of “seeds” and on different “crops.” As published on a recent poster from ASGCT-2022 authored by Mary Doolin, Ph.D., et al, [21] conditioning hMSCs with RoosterGEM culture during mRNA lipofection provides big jumps (from ~10% transfected to ~80%) in the proportion of mRNA (GFP+) transfected cells compared with Opti-MEM. Not only does culturing the cells in RoosterGEM during transfection help, but the reagent also improves efficiency by complexing the lipofection reagent+mRNA mix in RoosterGEM instead of Opti-MEM (see Figure 1 and 2). Further, we observe no signs of toxicity or cell death under these conditions.
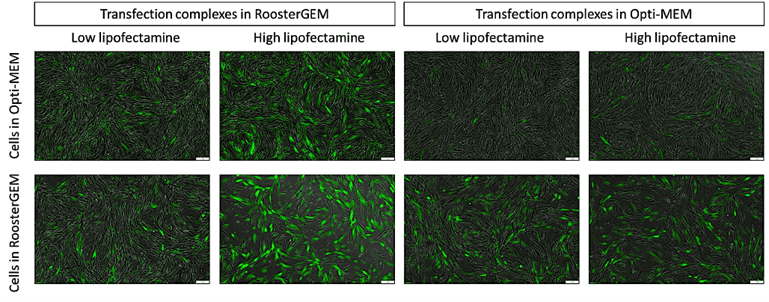
Figure 1, above. Fluorescence microscopy images of hMSCs transfected with GFP-mRNA using Lipofectamine3000 in RoosterGEM or Opti-MEM
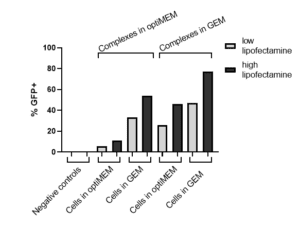 |
Figure 2, left. Quantified flow cytometry data indicating RoosterGEM as the optimal medium for cells and mRNA-lipid transfection complex formation. |
This product is designed for ease, so the user can instead focus on ID of best candidate mRNA performers to achieve therapeutic cellular “software upgrades.” [18] With efficiencies approaching 80%, zeroing in on the right mRNA design becomes a minimal effort! RoosterGEM’s intended use in mRNA transfection is as a drop-in replacement for MEM-type media in workflows that use lipofection or other chemical-based methods. This can be useful for “scale-out” into HTS for molecular discovery programs (example, Figure 3). As a complete medium, RoosterGEM may also be highly amenable to scale-up, with GMP product formats in development and near on the horizon. We’ll be writing on that topic soon with some interesting new findings to report. As shown, the experimental scheme is no more complex than your standard, familiar protocol. View a more detailed protocol here.
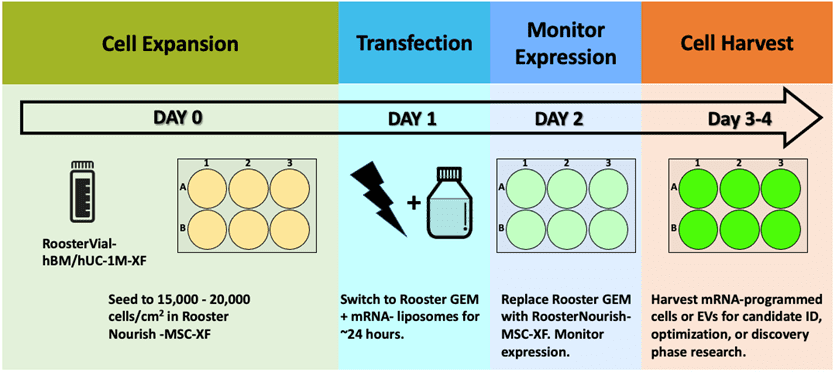
Figure 3, above. Example scheme and timeline for an mRNA transfection process used in R&D or candidate discovery research.
Long before RoosterGEM, nature evolved viruses to deliver and perpetuate genetic material across generations of progeny. But today’s nonviral modalities had to quickly recapitulate eons of natural selection out of little more than a bath (or catalog) of “primordial muck.” Yes, there’s been progress. Revisiting Professor Wolff’s 1998 analogy of nonviral gene delivery being stuck in the wild Cambrian Era, [1] it’s obvious that this realm of bioscience hasn’t yet felt a comparable “extinction event” to wipe the slate clean of most experimental configurations and usher in the Ordovician. Like the bizarre arthropods of old, one developmental “body plan” surely does not fit all, with diverse molecular technologies better suited for different tasks. In the case of ex vivo delivery of genetic material, we’ve found that a little cooperation (symbiosis?) works best, between existing nucleic acid carrier reagents and media precisely engineered to contain them.
On that note, here are some more developments to share!
References
- Wolff, J. A. and V. S. Trubetskoy, The Cambrian period of nonviral gene delivery. Nat Biotechnol, 1998. 16(5): p. 421-2. 10.1038/nbt0598-421
- Wolff, J. A., The “grand” problem of synthetic delivery. Nat Biotechnol, 2002. 20(8): p. 768-9. 10.1038/nbt0802-768
- Suzuki, Y. and H. Ishihara, Difference in the lipid nanoparticle technology employed in three approved siRNA (Patisiran) and mRNA (COVID-19 vaccine) drugs. Drug Metab Pharmacokinet, 2021. 41: p. 100424. 10.1016/j.dmpk.2021.100424
- Padda, I. S., A. U. Mahtani, and M. Parmar, Small Interfering RNA (siRNA) Based Therapy, in StatPearls. 2022: Treasure Island (FL).
- Kumar, P., A. Nagarajan, and P. D. Uchil, DNA Transfection Mediated by Cationic Lipid Reagents. Cold Spring Harb Protoc, 2019. 2019(3). 10.1101/pdb.prot095414
- Maurisse, R., et al., Comparative transfection of DNA into primary and transformed mammalian cells from different lineages. BMC Biotechnol, 2010. 10: p. 9. 10.1186/1472-6750-10-9
- Moradian, H., et al., mRNA Transfection-Induced Activation of Primary Human Monocytes and Macrophages: Dependence on Carrier System and Nucleotide Modification. Sci Rep, 2020. 10(1): p. 4181. 10.1038/s41598-020-60506-4
- Minnaert, A. K., et al., Strategies for controlling the innate immune activity of conventional and self-amplifying mRNA therapeutics: Getting the message across. Adv Drug Deliv Rev, 2021. 176: p. 113900. 10.1016/j.addr.2021.113900
- Hare, D., et al., The Importance of Physiologically Relevant Cell Lines for Studying Virus-Host Interactions. Viruses, 2016. 8(11). 10.3390/v8110297
- Kariko, K., et al., Generating the optimal mRNA for therapy: HPLC purification eliminates immune activation and improves translation of nucleoside-modified, protein-encoding mRNA. Nucleic Acids Res, 2011. 39(21): p. e142. 10.1093/nar/gkr695
- Sample, P. J., et al., Human 5′ UTR design and variant effect prediction from a massively parallel translation assay. Nat Biotechnol, 2019. 37(7): p. 803-809. 10.1038/s41587-019-0164-5
- Kim, S. C., et al., Modifications of mRNA vaccine structural elements for improving mRNA stability and translation efficiency. Mol Cell Toxicol, 2022. 18(1): p. 1-8. 10.1007/s13273-021-00171-4
- Bloom, K., F. van den Berg, and P. Arbuthnot, Self-amplifying RNA vaccines for infectious diseases. Gene Ther, 2021. 28(3-4): p. 117-129. 10.1038/s41434-020-00204-y
- Qin, S., et al., mRNA-based therapeutics: powerful and versatile tools to combat diseases. Signal Transduct Target Ther, 2022. 7(1): p. 166. 10.1038/s41392-022-01007-w
- Meacham, J. M., et al., Enhanced intracellular delivery via coordinated acoustically driven shear mechanoporation and electrophoretic insertion. Sci Rep, 2018. 8(1): p. 3727. 10.1038/s41598-018-22042-0
- Fu, Y., et al., Inhibition of cGAS-Mediated Interferon Response Facilitates Transgene Expression. iScience, 2020. 23(4): p. 101026. 10.1016/j.isci.2020.101026
- Takizaki, Misato, et al., Enhancing mechanism of gene transfection by heat shock. Chemistry Letters, 2017. 46(8): p. 1158-1160.
- Carson, J. and Candiello, J. RoosterGEM™: Addressing Key Challenges in Efficient Gene Transfer to Primary Cells. 2021; Available from: https://share.hsforms.com/1JFfgEWkRSVSxhfupTkjyMA3564o.
- Willstaedt, TM, et al., Development of an optimized lentiviral transduction medium and process to manufacture genetically modified MSC working cell banks. Cytotherapy, 2021. 23(5): p. S45.
- RoosterBio. RoosterBio Joins the Action at ISCT, ASGCT, & ISEV in May 2022. 2022; Available from: https://www.roosterbio.com/blog/roosterbio-joins-the-action-at-isct-asgct-isev-in-may-2022/.
- Doolin, M. T., T. M. Willstaedt, and J. A. Rowley, 138 – Mesenchymal Stem/Stromal Cells: OPTIMIZATION OF TRANSIENT TRANSFECTION OF MESENCHYMAL STROMAL CELLS WITH PLASMID DNA OR MESSENGER RNA. Cytotherapy, 2022. 24(5, Supplement): p. S59. https://doi.org/10.1016/S1465-3249(22)00196-7
